In collaboration with the Sharma Lab at Harvard Medical School, we study the impact of miRNAs and lincRNAs in the interactome to prioritize their role in diseases. We also built a disease similarity map based on shared miRNAs, showing high resolution to stratify disease subtypes.
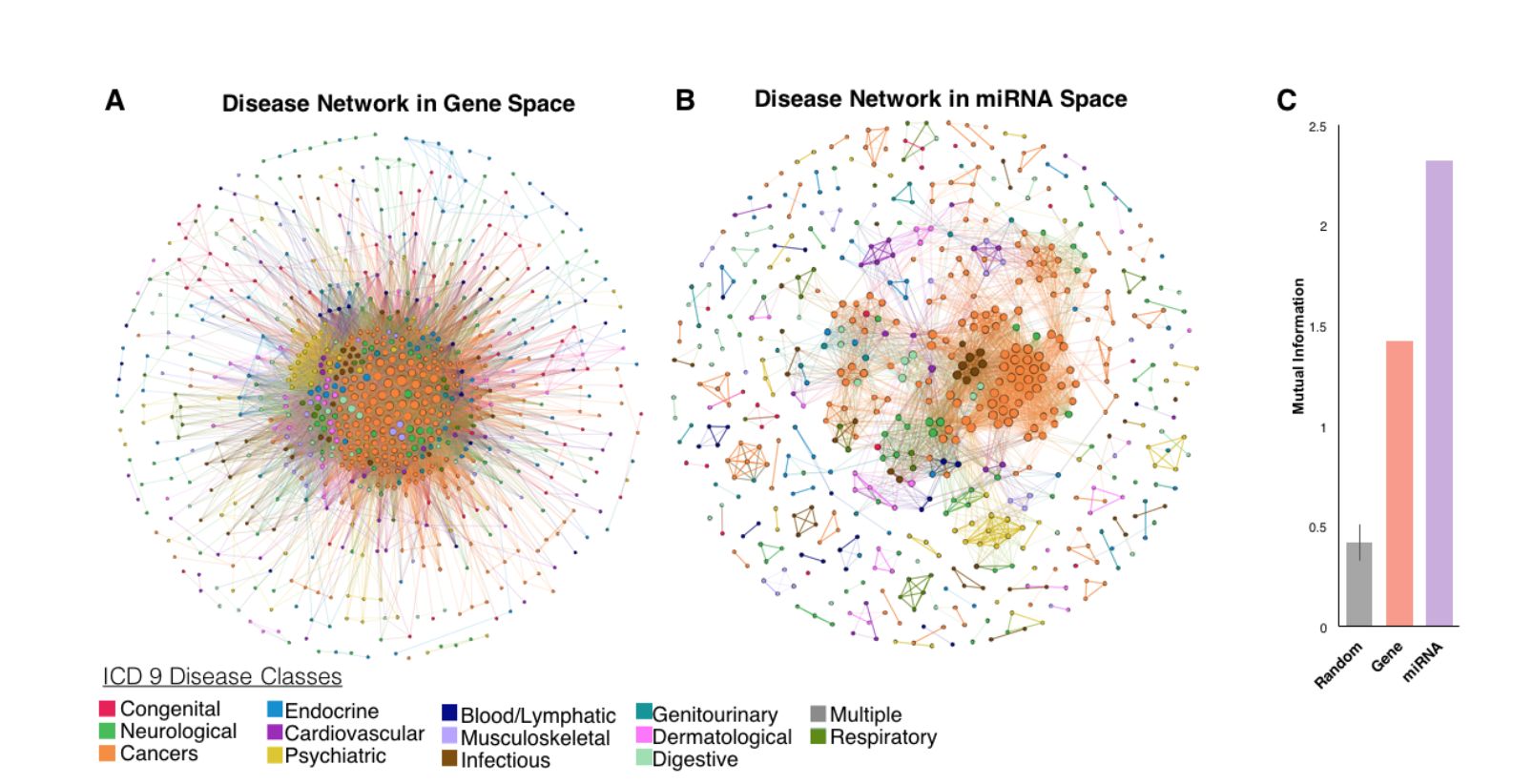
Abstract: Diseases are not driven by a single biological mediator but arise from perturbations in cellular interaction networks. As small non-coding RNAs that regulate expression of multiple disease related genes, miRNAs are part of a complex regulatory network where each miRNA regulates several genes and a gene is regulated by several miRNAs. While gene-disease networks and their emergent properties are well studied, the mapping of miRNA-disease networks is in its infancy. We take a conceptually different approach to prior miRNA-disease network studies. First, we predict a miRNA-disease network model without a priori information about miRNA-disease associations and validate predictions with experimental data. Second we explore how diseases and miRNAs are linked at a higher level of association through analysis of the miRNA-disease network’s emergent properties. We first construct a multipartite miRNA-gene-disease network by joining miRNA-gene, gene-gene, and gene-disease bipartite networks. We collapse the multipartite network with a network diffusion algorithm utilizing random walks that scores miRNA candidates by their proximity to disease genes, yielding the final weighted miRNA-disease network (MDN). The MDN is validated with experimental miRNA-disease data from dbDEMC; strong, significant positive correlations (r>0.25, p<0.0001, for six example cancers) between differential miRNA expression and our predicted miRNA-disease edge weights demonstrate biologically accurate miRNA rankings. We uncovered miRNA-based disease subtype classification from disease projections, which have been explored in gene space but not in miRNA space. We threshold the MDN to retain only high-ranked miRNAs per disease, then create a disease projection in miRNA space (DDM), where nodes are diseases and two diseases are connected if they share the same miRNA. Next, we collapse the diseases into broad disease types using ICD-9 medical codes. In comparison to a gene space disease projection, the DDM has greater biologically relevant disease clustering by disease type. The DDM has distinguishable, homogenous clusters, arising from phenotypically similar diseases being regulated by the same miRNAs. In contrast, the gene space projection is densely connected with fewer distinguishable clusters. We quantify this disease clustering by type using mutual information, which measures the alignment between structural communities and disease type. Mutual information in miRNA space is 2.3, compared to only 1.42 in gene space, indicating diseases tend to agglomerate by type in miRNA space more so than in gene space. Our miRNA-disease network model for disease subtyping enables more accurate and comprehensive disease-disease relationships and prioritization of disease-specific miRNAs critical to a disease subgroup.