Jule Timm interviewed me as part of her MA research at Aalto University on resonance in artistic practice. Starting from resonance singing — a collective vocal practice I have been developing for exploring emergence in groups — the conversation ranged across the physics of resonance, somatic practice, Tibetan dharma art, Balinese ritual theater, and the conditions under which art might be designed to truly resonate.
Exploring the role of care in educational systems
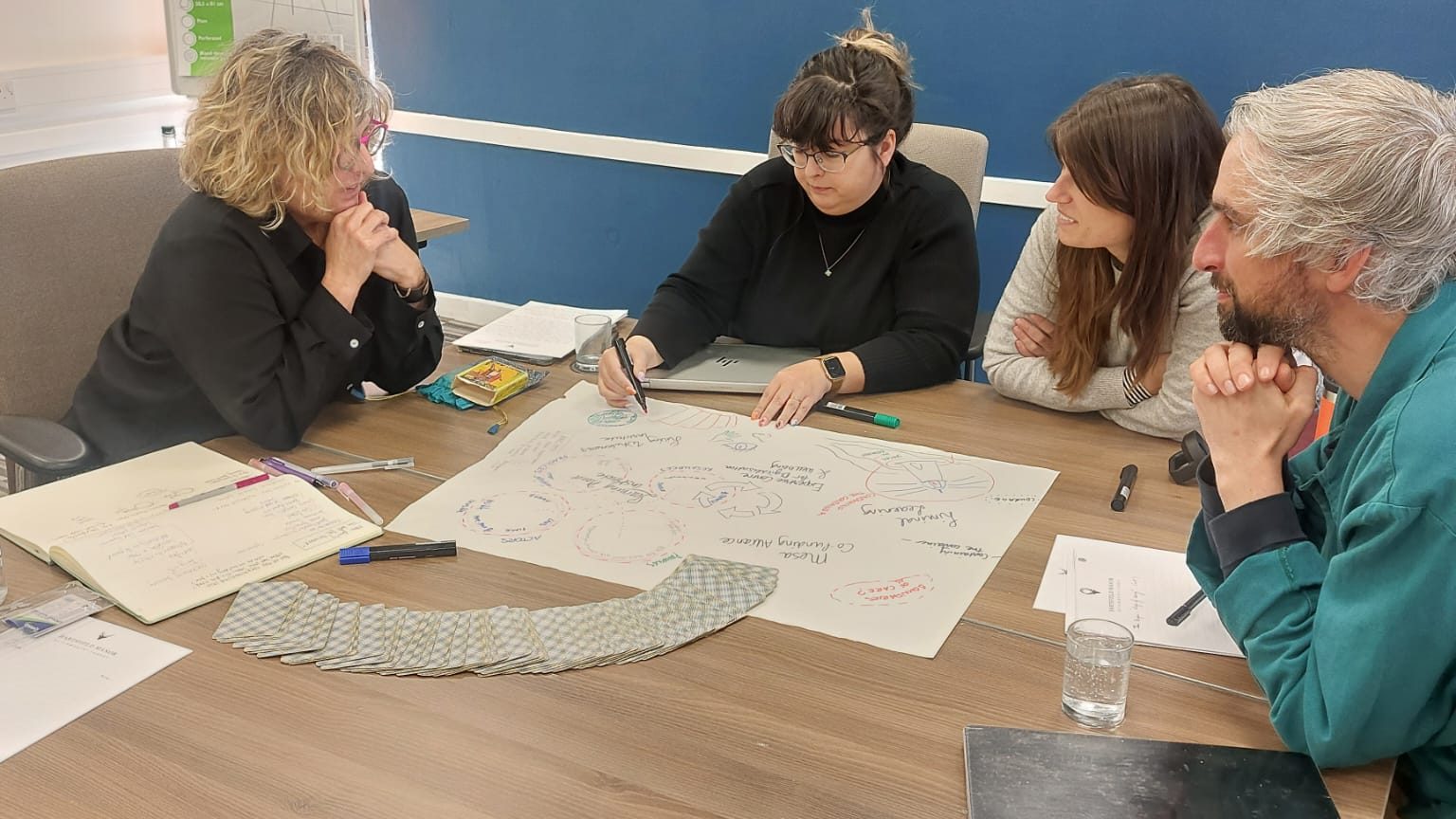

Over the past weeks, I found myself moving between two different and deeply connected spaces in the UK.
At the University of Cambridge, I gave a talk at the THRiVE research group on participatory and embodied approaches to learning. A few days later, I joined the Bloombox gathering, a small retreat bringing together educators, researchers, and theologians to explore care and the deeper purposes of education.
One space was structured around research, methods, and conceptual clarity. The other unfolded through dialogue, presence, and shared inquiry. Beneath these differences, both seemed to circle the same question:
How do we learn — and design systems — not only to solve problems, but to remain together in the face of difference, uncertainty, and conflict?
Continue readingIntegrating Behavior, Text, and Networks to Forecast Online Participation
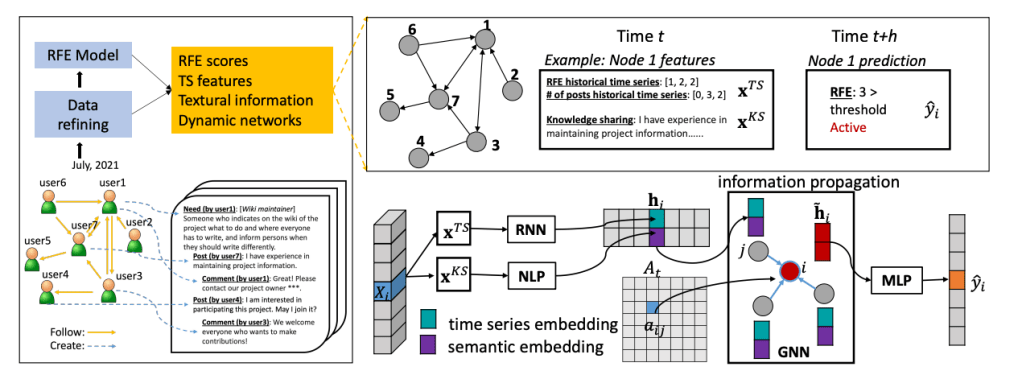
As online platforms increasingly rely on voluntary contributions—from open science to collaborative innovation—the ability to anticipate user engagement becomes both a scientific and practical priority. Yet predicting who will stay active, who will disengage, and why, remains a complex challenge. Our recent paper, KEGNN: Knowledge-Enhanced Graph Neural Networks for User Engagement Prediction (Fan et al., International Conference on Multimedia Retrieval 2025), introduces a novel framework that addresses this gap by integrating behavioral, social, and semantic signals into a unified predictive model.
Continue readingDrawing the Field: Art, Embodiment, and the Science of Group Dynamics
Over several days, I joined a group of approximately thirty educators, artists, and researchers for the Crafting Pedagogies of Togetherness residency—a prototype initiative investigating how embodied awareness practices can inform both educational pedagogy and collaborative methodologies. The residency was designed and facilitated by Studio Atelierista as part of a project co-funded by Erasmus+, and took place in a rural studio context, functioning as a site of transdisciplinary experimentation. Together, we articulated and tested new forms of learning that are relational, affectively attuned, and somatically grounded.
Continue readingThe Art & Science of Collective Intelligence: Podcast interview on R&D Unplugged
How can we foster collective intelligence in times of uncertainty, fragmentation, and crisis?
I had the pleasure of being interviewed on the R&D Unplugged Podcast by the Learning Planet Institute, where we explored how collaborative intelligence emerges—not just from technology or data, but from the quality of our relationships, processes, and ways of being together. We discussed how psychological safety, participatory methodologies, and even wisdom traditions can inform how we organize, learn, and navigate complexity—whether in research, education, or systemic change.
If you’re interested in how science, facilitation, and social healing intersect, I invite you to tune in:
🎙️ Listen to the episode
Relational Infrastructures for Collective Sensemaking and Action
How can we design containers that support relational development, foster collective intelligence, and cultivate the conditions for systemic transformation?
On March 20, I had the pleasure of presenting our work at the 11th edition of R&D Unplugged, hosted by the Research Unit Learning Transitions at the Learning Planet Institute. In this talk, I explored the intersection of network science, participatory methodologies, and wisdom traditions, highlighting how multi-level group practices and process-aware monitoring can help track and enhance relational quality over time.
Continue readingTeams That Thrive: AI-Driven Collaboration for Youth Participation
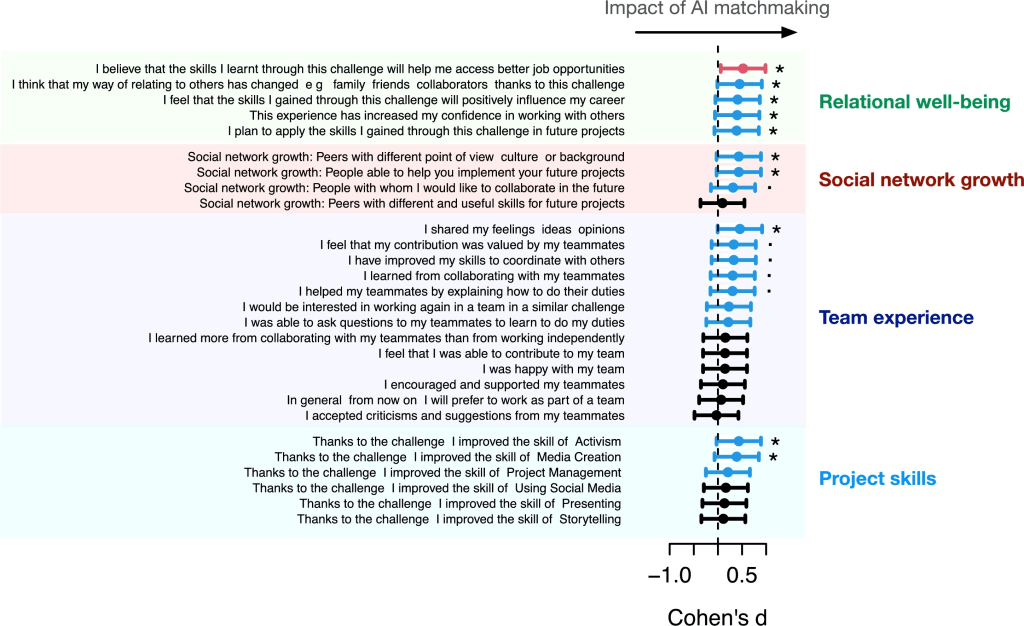
Team success isn’t just about outcomes—it’s also about how people feel, relate, and engage along the way. Understanding and improving this human dimension of participation is key to building teams that flourish. That’s the question we set out to answer in our latest study in the journal Computers and education: Artificial Intelligence, a collaboration between the Artificial Intelligence Research Institute in Spain and our team at the Learning Planet Institute in Paris. Together, we explored how artificial intelligence can help compose better teams in Challenge-Based Learning (CBL) environments, with a special focus on participants’ experiences—what we call participation quality.
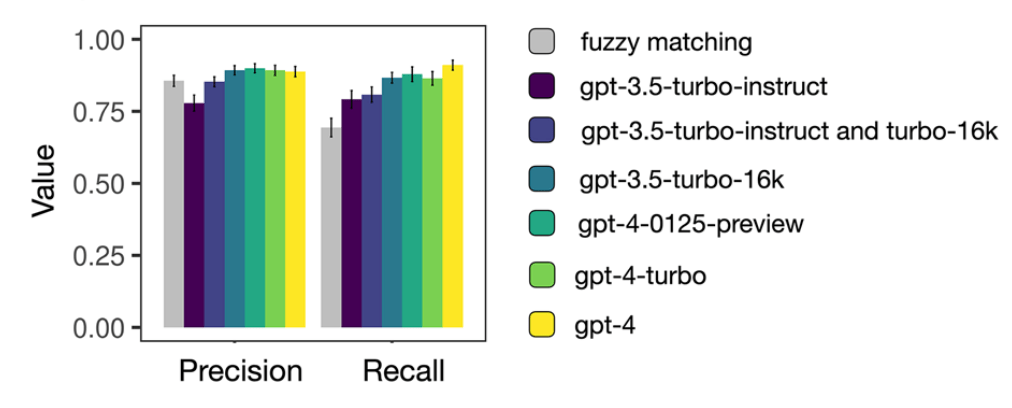
Continue readingFrom Text to Network: Mapping Scientific Collaboration Using LLMs
Understanding how scientists collaborate is key to improving research, but much of that collaboration is informal and buried in unstructured text. In our new article published in Applied Network Science, we show how Large Language Models (LLMs) can uncover these hidden networks—retrieving both inter-team collaborations and intra-team task allocations from free-form text with high accuracy.
Continue readingBuilding a Data-Driven Science of Citizen Science – My Reflections from ECSA 2024
At ECSA 2024, I shared my perspective on how data science can help us better understand and support participation in citizen science. I argued for a shift beyond traditional metrics like publications and citations, toward measuring the quality of engagement—using indicators such as relational well-being and collaborative dynamics.
Continue reading